Class: CHEM 0500 – Inorganic Chemistry
Instructor(s): Dr. Eric Victor
Student(s): Jon Mallen (Biophysics Sc.B. ’22), Yolanda Candler, Myung Joo Lee, Nicholas Moreno, Vivian Yuen
”My opinion about research at Brown has only been improved. Learning about bacteriophages in my first year and discovering my own, and now taking part in the Inorganic Chemistry CURE last semester, has confirmed my affinity for laboratory research, and also allowed me to better pinpoint my interests, skills, and weaknesses.”
–John Mallen
Description:
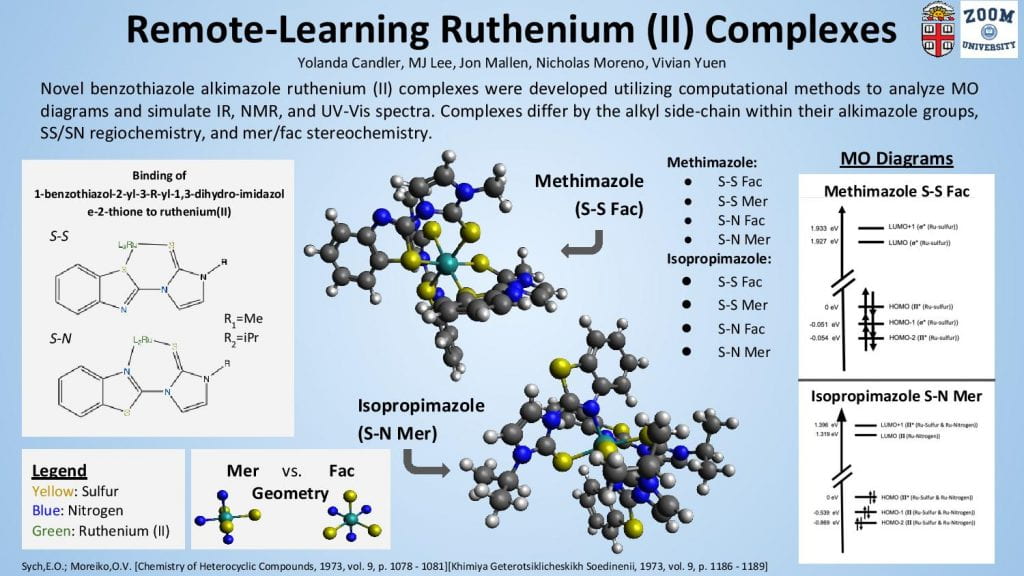
We computationally developed, optimized, and extracted energetic and physical parameters for the structures of eight novel benzothiazole alkimazole ruthenium (II) complexes through the Avogadro and ORCA software packages. Four complexes employed a benzothiazole methimazole ligand that was bound to the ruthenium (II) with either syn or anti sulfur-sulfur coordination in either the fac or mer geometry, and the other four employed a benzothiazole isopropimazole with the same syn/anti and fac/mer variation as its methimazole counterpart. There is no clear preference for coordination site and geometry among the different methimazole and isopropimazole in each of their four comparitive structures. However, the small difference in alkyl side-chain between the methimazole and isopropimazole has a significant effect on the resulting energetics of the involved complexes, with the isopropimazole variants uniformly more lower in energy than the methimazole complexes. This is evidence of greater trends in the complexation of ruthenium (II) with benzothiazole alkimazole ligands and is thus a part of the empirical data that will contribute to the future design of chemical routes employing this class of molecules.