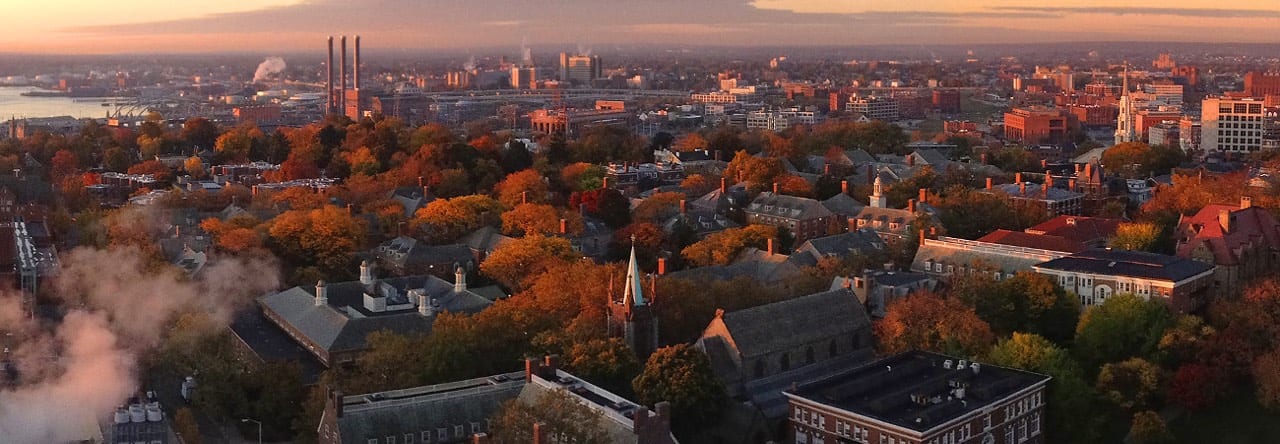
Current research in our group is focused on nanomechanics of engineering and biological systems.
For engineering systems, we study deformation, diffusion, growth, grain boundaries, stress evolution and failure in thin films, nanocrystalline materials, novel structural and functional materials, and energy materials.
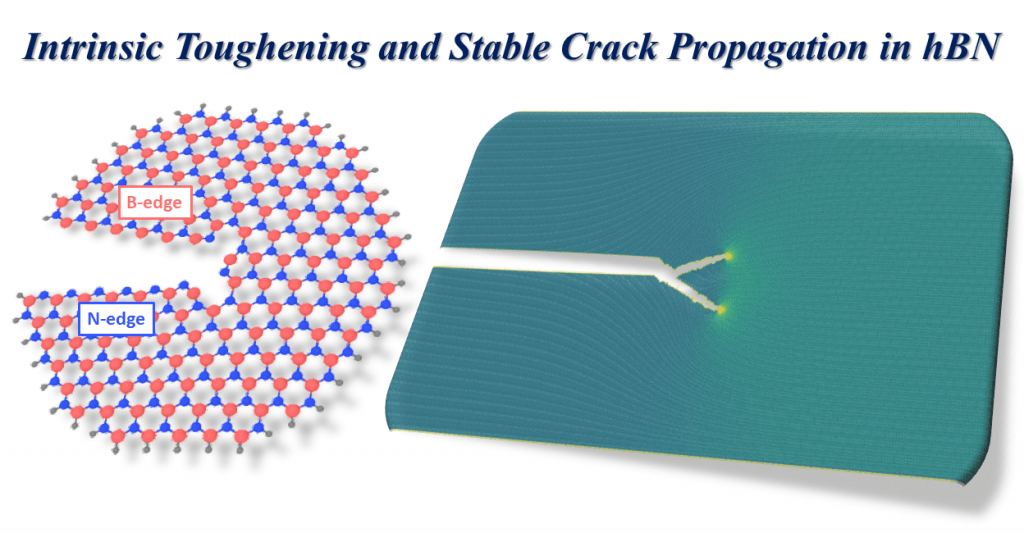
Our recent publication on Nature unveils an intrinsic toughening mechanism in h-BN, the toughest nanomaterial measured to date. (NTU press release)
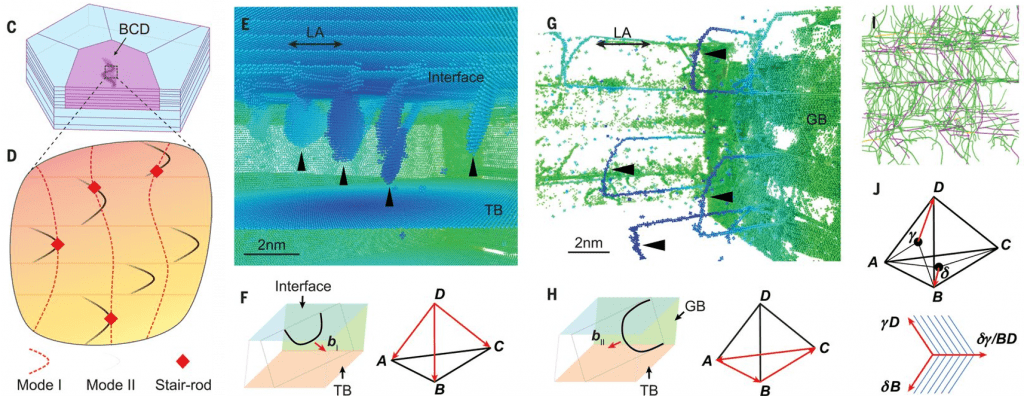
Our large-scale atomistic simulations revealed the mechanisms of GNT strengthening and associated deformation. (link to this paper)
For biomechanics and biological systems, we employ continuum mechanics, statistical mechanics and atomistic simulations to explore the ingenius mechanical principles from biological molecules, materials and systems such as peptides, proteins, cell membranes, bone, heart, and gecko, among others. The critical issues under investigation include stiffness, toughness, adhesion, size and shape effects, flaw tolerance, self-assembly, endocytosis, etc.
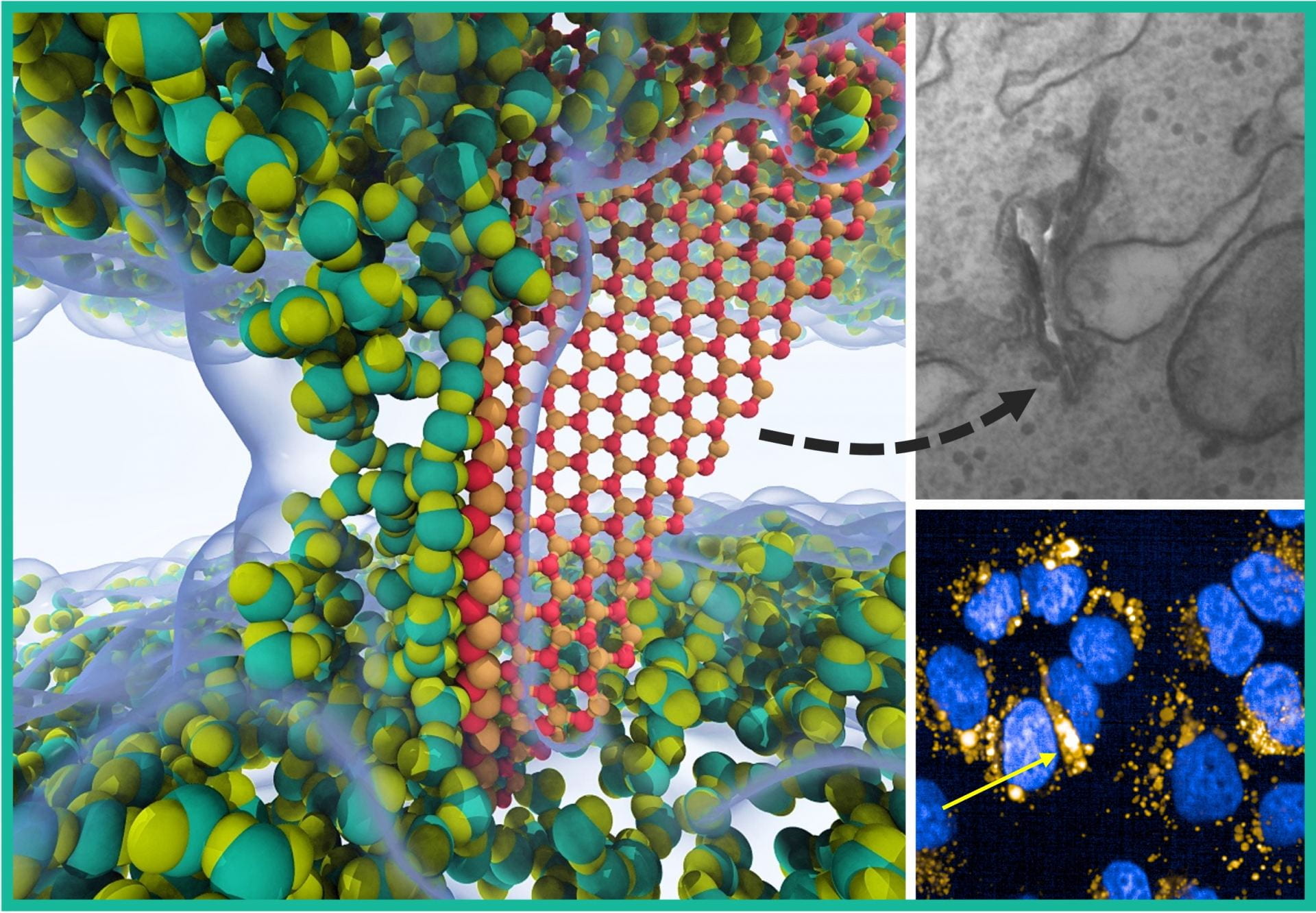
In our recent publication on Advanced Materials, we discovered for the first time a unique interaction mechanism between hBN and lipid membrane: a sharp hBN could penetrate a lipid bilayer and form a cross-membrane water channel along its exposed polar edges. This mechanism is fundamentally different from all known membrane damaging mechanisms of 2D materials and may have general implications for all binary or multielement 2D materials with polar edges.
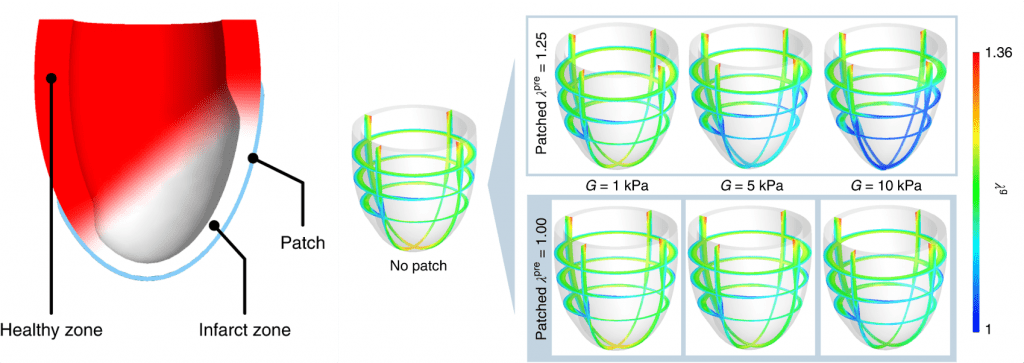
Left: illustration of the patched heart model with infarct zone. Right: distribution of the lengthening factor after dilatation under various values of patch stiffness and pre-stretch. (link to this paper)
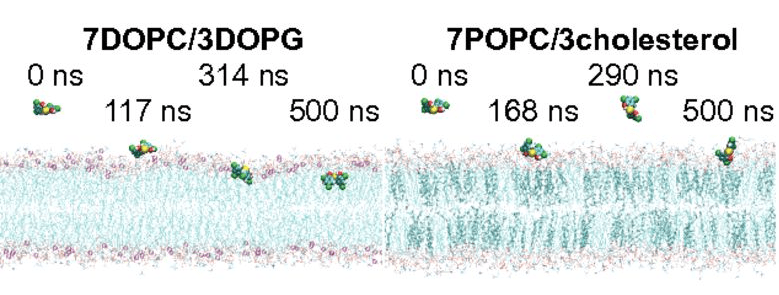
MD simulations show that Bithionol, a clinically approved anthelmintic drug, kills MRSA persisters by selectively disrupting membrane lipid bilayers. (link to this paper)
